AxonEM Dataset: 3D Axon Instance Segmentation of Brain Cortical Regions
6Universidade do Vale do Rio dos Sinos 7Argonne National Laboratory 8Donostia International Physics Center (DPIC)
9University of the Basque Country (UPV/EHU) 10Ikerbasque, Basque Foundation for Science
[Paper ] [Code] [Dataset]

Abstract
Electron microscopy (EM) enables the reconstruction of neural circuits at the level of individual synapses, which has been transformative for scientific discoveries. However, due to the complex morphology, an accurate reconstruction of cortical axons has become a major challenge. Worse still, there is no publicly available large-scale EM dataset from the cortex that provides dense ground truth segmentation for axons, making it difficult to develop and evaluate large-scale axon reconstruction methods. To address this, we introduce the AxonEM dataset, which consists of two 30x30x30 μm3 EM image volumes from the human and mouse cortex, respectively. We thoroughly proofread over 18,000 axon instances to provide dense 3D axon instance segmentation, enabling large-scale evaluation of axon reconstruction methods. In addition, we densely annotate nine ground truth subvolumes for training, per each data volume. With this, we reproduce two published state-of-the-art methods and provide their evaluation results as a baseline. We publicly release our code and data here to foster the development of advanced methods.
Dataset
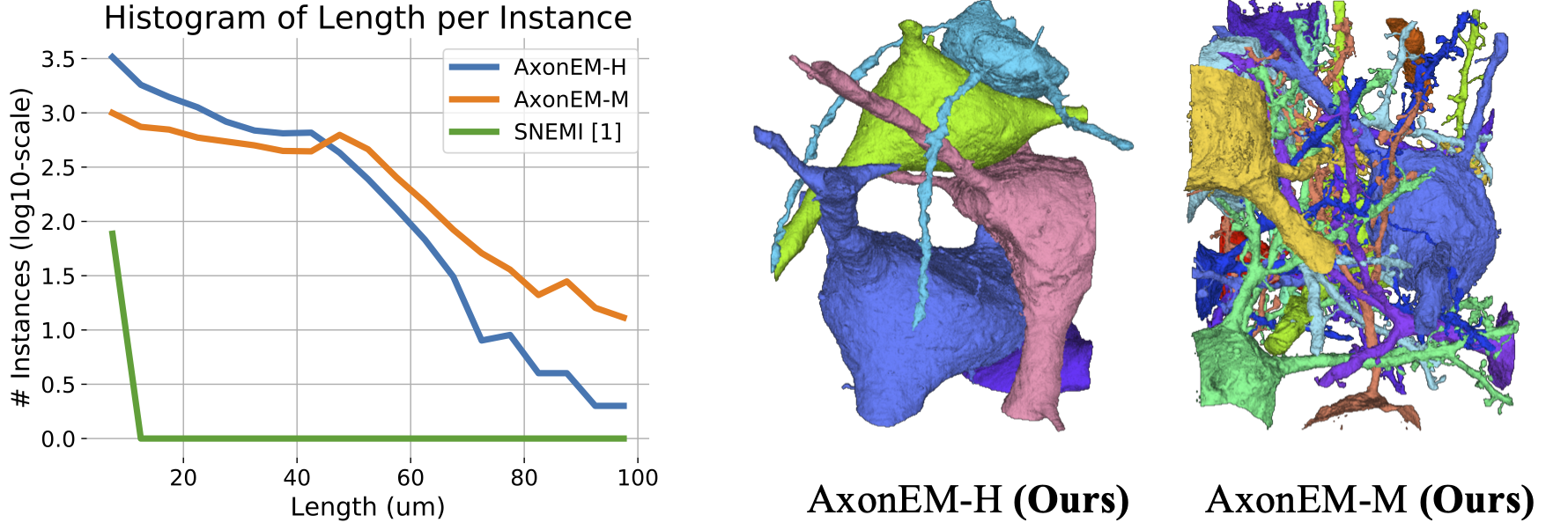
(Left) We plot the distribution of the axon instance length for AxonEM and SNEMI3D datasets. (Right) We show the 3D rendering of neurons with somas in AxonEM dataset. Two large glial cells (not rendered) in AxonEM-H occupy the space between the blue and pink neurons, leading to much fewer long axons compared to AxonEM-M.
Citation
@misc{wei2021axonem,
title={AxonEM Dataset: 3D Axon Instance Segmentation of Brain Cortical Regions},
author={Donglai Wei and Kisuk Lee and Hanyu Li and Ran Lu and J. Alexander Bae and Zequan Liu and
Lifu Zhang and Márcia dos Santos and Zudi Lin and Thomas Uram and Xueying Wang and Ignacio
Arganda-Carreras and Brian Matejek and Narayanan Kasthuri and Jeff Lichtman and Hanspeter Pfister},
year={2021},
eprint={2107.05451},
archivePrefix={arXiv},
primaryClass={cs.CV}
}
Acknowledgement
KL, RL, and JAB were supported by the Intelligence Advanced Research Projects Activity (IARPA) via Department of Interior/ Interior Business Center (DoI/IBC) contract number D16PC0005, NIH/NIMH (U01MH114824, U01MH117072, RF1 MH117815), NIH/NINDS (U19NS104648, R01NS104926), NIH/NEI (R01EY027 036), and ARO (W911NF-12-1-0594), and are also grateful for assistance from Google, Amazon, and Intel. DW, ZL, JL, and HP were partially supported by NSF award IIS-1835231. I. A-C would like to acknowledge the support of the Beca Leonardo a Investigadores y Creadores Culturales 2020 de la Fundación BBVA. We thank Viren Jain, Michał Januszewski and their team for generating the initial segmentation for AxonEM-H, and Daniel Franco-Barranco for setting up the challenge using AxonEM.
Declaration of Interests
KL, RL, and JAB disclose financial interests in Zetta AI LLC.